SPPARKS Kinetic Monte Carlo Simulator
Anyone who considers arithmetical methods of producing random digits is,
of course, in a state of sin. -- John von Neumann
God does not play dice -- Albert Einstein
This is the home page for the kinetic Monte Carlo code SPPARKS, an
acronym for Stochastic Parallel PARticle Kinetic Simulator.
SPPARKS is a parallel Monte Carlo code for on-lattice and off-lattice
models that includes algorithms for kinetic Monte Carlo (KMC),
rejection kinetic Monte Carlo (rKMC), and Metropolis Monte Carlo
(MMC). It implements several KMC solvers whose serial computational
complexity ranges from O(N) to O(NlogN) to O(1) in the number of
events N owned by a processor. In a generic sense the solvers catalog
a list of "events", each with an associated probability, choose a
single event to perform, and advance time by the correct amount.
Events may be chosen individually at random, or by sweeping over sites
in a more ordered fashion.
Applications that are packaged with SPPARKS, or which a user adds to
the code, define sites and events and probabilities, as well as how
the execution of an event changes the state of the model.
SPPARKS runs on single processors or in parallel using message-passing
techniques and a spatial-decomposition of the simulation domain. The
code is designed to be easy to modify or extend with new
functionality.
SPPARKS is distributed as an open source code under
the terms of the GPL, or sometimes by request under the terms of
the GNU Lesser General Public License (LGPL). The current
version can be downloaded here.
SPPARKS was developed at Sandia National Laboratories, a US
Department of Energy laboratory. The Authors
page lists the developers and their contact info, along
with others who have contributed code to SPPARKS. Funding for SPPARKS
development has come from the DOE and is also acknowledged on the
Authors page.
Recent SPPARKS News
 (10/25) New HCP Burgers orientation variants and
Stitch api update. See full description of the 08Oct25 release on
this page.
(10/25) New HCP Burgers orientation variants and
Stitch api update. See full description of the 08Oct25 release on
this page.
 (11/24) New potts/quaternion application
to include relative lattice orientation differences between sites in
the Hamiltonian energy function which drives grain growth and
texturing. See full description of the 27Nov24 release on this
page.
(11/24) New potts/quaternion application
to include relative lattice orientation differences between sites in
the Hamiltonian energy function which drives grain growth and
texturing. See full description of the 27Nov24 release on this
page.
 (9/23) New potts/am/bezier application
for modeing additive manufacturing of metals, using Bezier curves to
fit the meltpool shape. See the description of the 6Sep23
release on this page.
(9/23) New potts/am/bezier application
for modeing additive manufacturing of metals, using Bezier curves to
fit the meltpool shape. See the description of the 6Sep23
release on this page.
 (6/22) New diffusion/multiphase
application which allows any number of different phases (atomic
species for an atomic-scale model) to be used, any of which can be
immobile (pinned) or mobile (diffusing). See the description of the
24Jun21 release on this page.
(6/22) New diffusion/multiphase
application which allows any number of different phases (atomic
species for an atomic-scale model) to be used, any of which can be
immobile (pinned) or mobile (diffusing). See the description of the
24Jun21 release on this page.
 (7/20) A variety of options were added
for modeling additive manufacturing processes via Potts model
simulations which evolve microstructure in the presence of a moving
laser spot which heats material. See the description of the 23Jul20
release on this page.
(7/20) A variety of options were added
for modeling additive manufacturing processes via Potts model
simulations which evolve microstructure in the presence of a moving
laser spot which heats material. See the description of the 23Jul20
release on this page.
 (7/20) A new I/O system called the Stitch
library has been added to SPPARKS. It was specially developed for
additive manufacturing (AM) simulations. Using Stitch, an output
database for microstructure simulations is created incrementally by
appending lattice sites much in the same way new material is added to
an AM part. See the dump stitch and set
stitch commands for details, as well as the stitch and
am_path folders in the examples directory.
(7/20) A new I/O system called the Stitch
library has been added to SPPARKS. It was specially developed for
additive manufacturing (AM) simulations. Using Stitch, an output
database for microstructure simulations is created incrementally by
appending lattice sites much in the same way new material is added to
an AM part. See the dump stitch and set
stitch commands for details, as well as the stitch and
am_path folders in the examples directory.
 (11/16) A variety of new input/output
options were added, including dump sites and dump
vtk styles, and dump_modify and
read_sites options. These make it easier to use
SPPARKS with VTK-compatible visualizers, like
ParaView, and to continue/restart SPPARKS
simulations using dump file snapshots as a restart file.
(11/16) A variety of new input/output
options were added, including dump sites and dump
vtk styles, and dump_modify and
read_sites options. These make it easier to use
SPPARKS with VTK-compatible visualizers, like
ParaView, and to continue/restart SPPARKS
simulations using dump file snapshots as a restart file.
 (3/12) Release of Python wrapper for
SPPARKS in python dir with additional examples of using SPPARKS as a
library in the example/COUPLE dir.
(3/12) Release of Python wrapper for
SPPARKS in python dir with additional examples of using SPPARKS as a
library in the example/COUPLE dir.
 (11/09) Release of 11 Nov 2009 version of
SPPARKS. New commands and features include: app_style
diffusion, app_style
relax, barrier,
create_box,
create_sites,
deposition, diag_style
propensity,
dimension,
dump_modify,
dump_one,
lattice
pair_coeff,
pair_style,
processors,
read_sites,
region, set,
undump, and Schwoebel hops via the
app_style diffusion command. See
details here.
(11/09) Release of 11 Nov 2009 version of
SPPARKS. New commands and features include: app_style
diffusion, app_style
relax, barrier,
create_box,
create_sites,
deposition, diag_style
propensity,
dimension,
dump_modify,
dump_one,
lattice
pair_coeff,
pair_style,
processors,
read_sites,
region, set,
undump, and Schwoebel hops via the
app_style diffusion command. See
details here.
 (9/09) First off-lattice parallel
application added.
(9/09) First off-lattice parallel
application added.
 (4/09) Initial open-source release of
SPPARKS.
(4/09) Initial open-source release of
SPPARKS.
SPPARKS Highlight
(see the Pictures & Movies page for more examples of
SPPARKS calculations)
This is work by Liz Holm (eaholm at sandia.gov) at Sandia and Todd
Hoffmann (todd.hoffmann at asu.edu) and his advisor at Carnegie
Mellon, to study the mechanisms and statistics associated with
abnormal grain growth. In grain growth, when the system is annealed,
grain boundaries will move and rearrange so as to increase the average
grain size and decrease the grain boundary area per unit volume. In
normal grain growth, this occurs relatively uniformly, but in abnormal
grain growth, one grain grows at the expense of the others. Abnormal
grain growth can have many causes. Here the effects of static particle
dispersions on grain growth are studied. Particles should pin grain
boundaries in positions that maximize their contact with the
particles, so that grain growth cannot proceed. However, occasionally
one grain breaks free and is able to grow at the expense of others,
leading to abnormal growth.
The images and movies show typical results for a 300x300x300 lattice,
run for a million Monte Carlo steps (sweep over all lattice sites),
with 5 volume percent pinning particles, inserted when the average
grain radius was 10 sites. In the normal case, the system becomes
fully pinned. In the second case, abnormal growth eventually occurs.
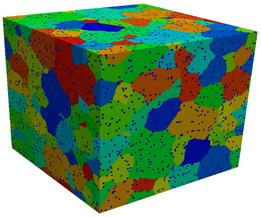
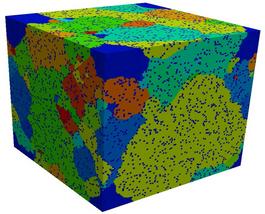


Two images and two 1.5 Mb AVI movies
 (10/25) New HCP Burgers orientation variants and
Stitch api update. See full description of the 08Oct25 release on
this page.
(10/25) New HCP Burgers orientation variants and
Stitch api update. See full description of the 08Oct25 release on
this page.
 (11/24) New potts/quaternion application
to include relative lattice orientation differences between sites in
the Hamiltonian energy function which drives grain growth and
texturing. See full description of the 27Nov24 release on this
page.
(11/24) New potts/quaternion application
to include relative lattice orientation differences between sites in
the Hamiltonian energy function which drives grain growth and
texturing. See full description of the 27Nov24 release on this
page.
 (9/23) New potts/am/bezier application
for modeing additive manufacturing of metals, using Bezier curves to
fit the meltpool shape. See the description of the 6Sep23
release on this page.
(9/23) New potts/am/bezier application
for modeing additive manufacturing of metals, using Bezier curves to
fit the meltpool shape. See the description of the 6Sep23
release on this page.
 (6/22) New diffusion/multiphase
application which allows any number of different phases (atomic
species for an atomic-scale model) to be used, any of which can be
immobile (pinned) or mobile (diffusing). See the description of the
24Jun21 release on this page.
(6/22) New diffusion/multiphase
application which allows any number of different phases (atomic
species for an atomic-scale model) to be used, any of which can be
immobile (pinned) or mobile (diffusing). See the description of the
24Jun21 release on this page.
 (7/20) A variety of options were added
for modeling additive manufacturing processes via Potts model
simulations which evolve microstructure in the presence of a moving
laser spot which heats material. See the description of the 23Jul20
release on this page.
(7/20) A variety of options were added
for modeling additive manufacturing processes via Potts model
simulations which evolve microstructure in the presence of a moving
laser spot which heats material. See the description of the 23Jul20
release on this page.
 (7/20) A new I/O system called the Stitch
library has been added to SPPARKS. It was specially developed for
additive manufacturing (AM) simulations. Using Stitch, an output
database for microstructure simulations is created incrementally by
appending lattice sites much in the same way new material is added to
an AM part. See the dump stitch and set
stitch commands for details, as well as the stitch and
am_path folders in the examples directory.
(7/20) A new I/O system called the Stitch
library has been added to SPPARKS. It was specially developed for
additive manufacturing (AM) simulations. Using Stitch, an output
database for microstructure simulations is created incrementally by
appending lattice sites much in the same way new material is added to
an AM part. See the dump stitch and set
stitch commands for details, as well as the stitch and
am_path folders in the examples directory.
 (11/16) A variety of new input/output
options were added, including dump sites and dump
vtk styles, and dump_modify and
read_sites options. These make it easier to use
SPPARKS with VTK-compatible visualizers, like
ParaView, and to continue/restart SPPARKS
simulations using dump file snapshots as a restart file.
(11/16) A variety of new input/output
options were added, including dump sites and dump
vtk styles, and dump_modify and
read_sites options. These make it easier to use
SPPARKS with VTK-compatible visualizers, like
ParaView, and to continue/restart SPPARKS
simulations using dump file snapshots as a restart file.
 (3/12) Release of Python wrapper for
SPPARKS in python dir with additional examples of using SPPARKS as a
library in the example/COUPLE dir.
(3/12) Release of Python wrapper for
SPPARKS in python dir with additional examples of using SPPARKS as a
library in the example/COUPLE dir.
 (11/09) Release of 11 Nov 2009 version of
SPPARKS. New commands and features include: app_style
diffusion, app_style
relax, barrier,
create_box,
create_sites,
deposition, diag_style
propensity,
dimension,
dump_modify,
dump_one,
lattice
pair_coeff,
pair_style,
processors,
read_sites,
region, set,
undump, and Schwoebel hops via the
app_style diffusion command. See
details here.
(11/09) Release of 11 Nov 2009 version of
SPPARKS. New commands and features include: app_style
diffusion, app_style
relax, barrier,
create_box,
create_sites,
deposition, diag_style
propensity,
dimension,
dump_modify,
dump_one,
lattice
pair_coeff,
pair_style,
processors,
read_sites,
region, set,
undump, and Schwoebel hops via the
app_style diffusion command. See
details here.
 (9/09) First off-lattice parallel
application added.
(9/09) First off-lattice parallel
application added.
 (4/09) Initial open-source release of
SPPARKS.
(4/09) Initial open-source release of
SPPARKS.

